Reem Ali Alghamdi
PhD student, Computer Science
Publications
Research Experience
-
2022 - Present
Graduate Student Researcher
King Abdullah University of Science and Technology, Saudi Arabia
- Advisor: Professor Markus Hadwiger
- Reseach focus: Visualization in Astrophysics, High Performance Computing, Scientific Visualization, GPU Programming
-
2024 - 2024
Visiting Researcher
Linköping University, Norrköping, Sweden
-
2021 - 2021
Graduate Student Researcher
King Abdullah University of Science and Technology, Saudi Arabia
- Advisor: Professor Xiangliang Zhang
- Reseach focus: Deep Learning, Natural Language Processing, Transfer Learning, Math Word Problems
Mentoring And Teaching Experience
-
2025 - 2025
Teaching Assistant - KAUST MEngg in AI
Ministry of Interior, Saudi Arabia
- I worked as a TA for The Applied Data Analytics course
-
2021 - 2025
Graduate Courses Teaching Assistant
King Abdullah University of Science and Technology, Saudi Arabia
- CS380 - GPU and GPGPU Programming
- CS247 - Scientific Visualization
- CS220 - Data Analytics
- CS207 - Programming Methodology and Abstractions
- CS201 - Introduction to Programming with Python
- I worked as a TA for multiple courses. Responsibilities included writing homeworks and tests, grading, recitation sessions, one-on-one student support
-
2025 - 2025
Instructor - AI Specialization
KAUST Academy, Saudi Arabia
- Introduction to Artificial Intelligence: Taught students machine learning concepts such as linear regression, logistic regression and an introduction to deep learning.
-
2022 - 2023
Teaching Assistant - KAUST MS in Data Science and Analytics
ARAMCO, Saudi Arabia
- Information Visualization and Visual Analytics: InfoVis and Visual Analytics course and an overview of techniques and tools for interactive and animated visualization, including Javascript, D3.js, Vega-Lite, Vega-Altair, observable
- Practical Tools for Machine Learning: An introduction to Python programming language course and relevant libraries for data science (numpy, pandas, matplotlib)
-
Aug 2021
Python High School Mentor For KAUST Virtual Academy Program
King Abdullah University of Science and Technology, Saudi Arabia
Conferences And Presentations
-
Jun 2023
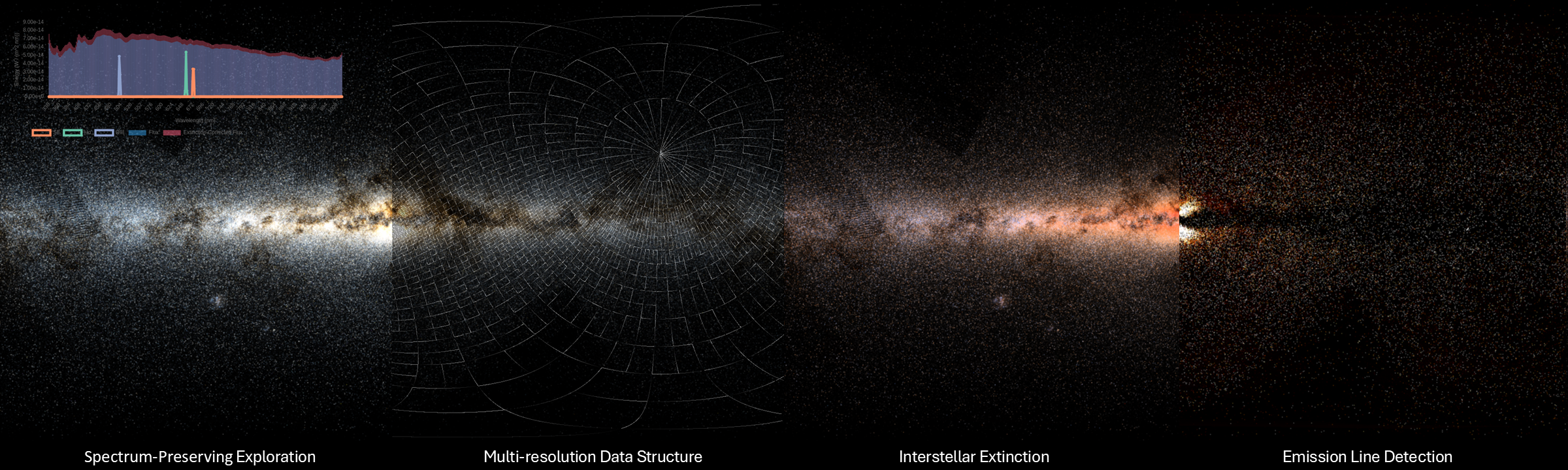
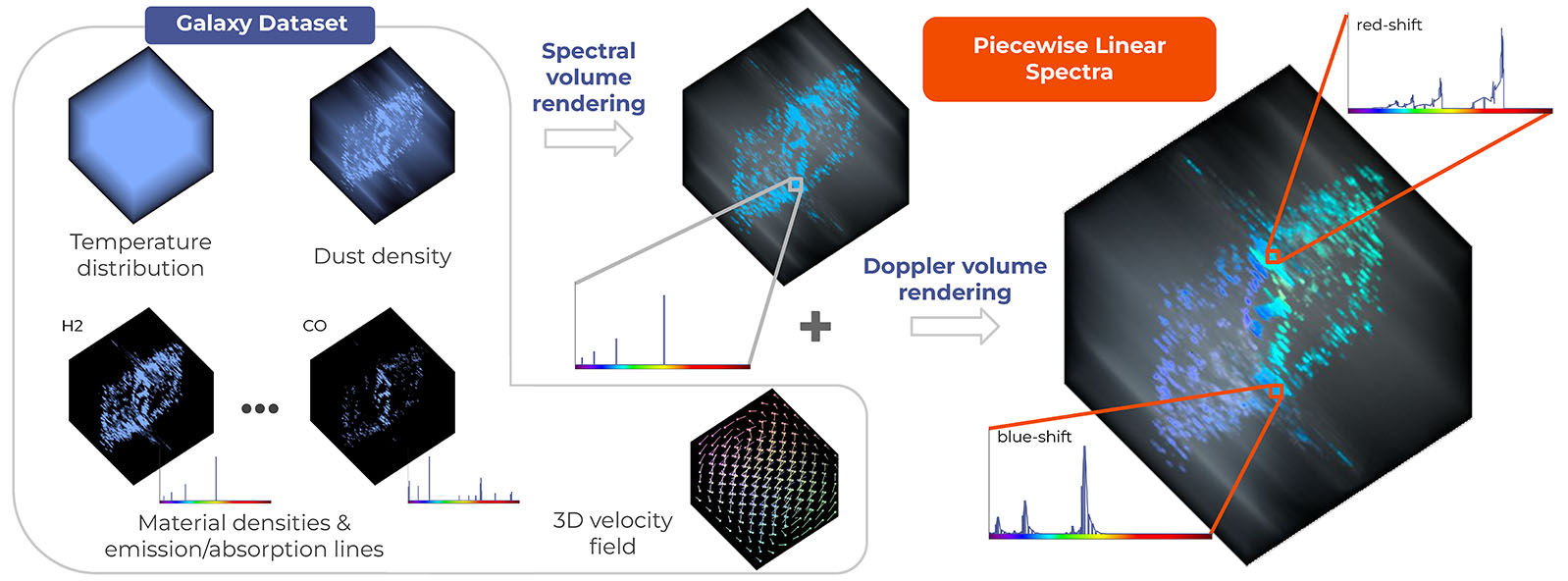
Doppler Volume Rendering
The 25th edition of EuroVis, Leipzig, Germany
- I presented our paper Doppler Volume Rendering at EuroVis 2023 conference
-
Jun 2022
Education
-
2022 - Present
PhD, Computer Science
King Abdullah University of Science and Technology, Saudi Arabia
- Advisor: Professor Markus Hadwiger
- Focus: Visualization in Astrophysics, High Performance Computing, Scientific visualization, GPU programming
-
2020 - 2021
Master's Degree, Computer Science
King Abdullah University of Science and Technology, Saudi Arabia
- Thesis Advisor: Professor Xiangliang Zhang
- Thesis: Solving Arabic Math Word Problems via Deep Learning
- Focus: AI and Scientific Visualization
- I Took courses in Scientific Visualization, GPU programming, AI, Machine Learning, Deep Learning, Computer Vision, Reinforcement Learning, Bioinformatics, Numerical Linear Algebra, and Probability & Statistics in R
-
2016 - 2020
Bachelor's Degree, Computer Science
Princess Nourah Bint Abdulrahman University, Saudi Arabia
Technical And Soft Skills
Programming Languages
Python
C/C++
CUDA
JavaScript
HTML
CSS
MATLAB
Java
C#
R
SQL
Bash
Tools
OpenGL
Qt
PyTorch
Numpy
Pandas
OpenCV
Flask
ReactJS
JQuery
D3
Android
Linux
Technical Skills
GPU Programming
Computer Graphics
Visualization
Data Analysis
Artificial Intelligence
Deep Learning
Software Development
Servers
Soft Skills
Leadership And Team Work
Time Management And Planning
Initiative And Self-motivation
Problem-Solving And Creativity
Working Under Pressure
Communication And Interpersonal Skills
Volunteering
English, Arabic And Japanese